Tracking options#
We explore the impact of some of the tracking options for two of the sample datasets.
import warnings
from string import ascii_lowercase
import cartopy
import cartopy.crs as ccrs
import matplotlib as mpl
import matplotlib.patheffects as path_effects
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import xarray as xr
from IPython.display import display, Markdown
from joblib import delayed, Parallel
import tams
warnings.filterwarnings("ignore", category=cartopy.io.DownloadWarning)
plt.rcParams.update(
{
"axes.formatter.use_mathtext": True,
}
)
%matplotlib inline
xr.set_options(display_expand_data=False)
Show code cell output
<xarray.core.options.set_options at 0x230061013f0>
For sample satellite data#
tb = tams.load_example_tb()
tb
<xarray.DataArray 'ch9' (time: 6, y: 714, x: 2829)>
295.9 295.9 295.8 295.8 295.8 295.9 295.9 295.8 ... nan nan nan nan nan nan nan
Coordinates:
lon (time, y, x) float32 ...
lat (time, y, x) float32 ...
* time (time) datetime64[ns] 2006-09-01 ... 2006-09-01T10:00:00
Dimensions without coordinates: y, x
Attributes:
units: K
long_name: Brightness temperatureIdentify cloud elements (CEs)#
Our tracking options cases here are all in the tracking stage, so we only have to run tams.identify() once.
%%time
ces, _ = tams.identify(tb, parallel=True)
Show code cell output
[Parallel(n_jobs=-2)]: Using backend LokyBackend with 11 concurrent workers.
CPU times: total: 172 ms
Wall time: 5.26 s
[Parallel(n_jobs=-2)]: Done 2 out of 6 | elapsed: 5.0s remaining: 10.1s
[Parallel(n_jobs=-2)]: Done 3 out of 6 | elapsed: 5.0s remaining: 5.0s
[Parallel(n_jobs=-2)]: Done 4 out of 6 | elapsed: 5.0s remaining: 2.5s
[Parallel(n_jobs=-2)]: Done 6 out of 6 | elapsed: 5.1s finished
Run cases#
%%time
cases = {}
time = tb.time.values.tolist()
proj = -15
with warnings.catch_warnings():
warnings.filterwarnings("ignore", message="forward `look` considered experimental")
cases["default"] = tams.track(ces, time)
cases["link=f, norm=min"] = tams.track(ces, time, look="f", overlap_norm="min")
cases["link=f, norm=max"] = tams.track(ces, time, look="f", overlap_norm="max")
cases[f"u_proj={proj}"] = tams.track(ces, time, u_projection=proj)
cases[f"u_proj={proj}, link=f, norm=min"] = tams.track(ces, time, u_projection=proj, look="f", overlap_norm="min")
cases[f"u_proj={proj}, link=f, norm=max"] = tams.track(ces, time, u_projection=proj, look="f", overlap_norm="max")
for key, ce in cases.items():
cases[key] = tams.classify(ce)
CPU times: total: 5.83 s
Wall time: 5.83 s
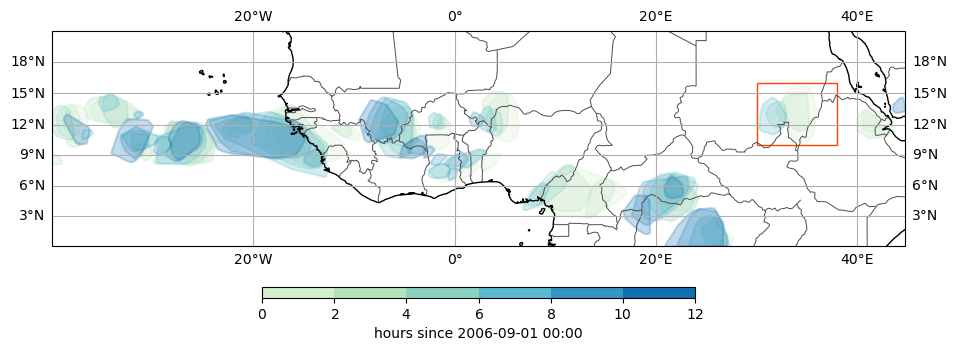
Plot CEs#
And check if the method produced a single track within the box, saving a table of these results.
m, n = 2, 3
assert len(cases) <= m * n
# Restrict range
# lon_min, lat_min, lon_max, lat_max = -10, 8, -3, 15
# lon_min, lat_min, lon_max, lat_max = 17, 1, 27, 8
# lon_min, lat_min, lon_max, lat_max = 20, 1, 27, 4
lon_min, lat_min, lon_max, lat_max = 30, 10, 38, 16
cases_full = []
for key in cases:
d = {"u_proj": 0, "link": "b", "norm": "a"}
updates = {}
if key != "default":
for kv in key.split(", "):
k, v = kv.split("=")
updates[k] = v
if "norm" not in updates and updates.get("link", "b").startswith("f"):
d["norm"] = "b"
d.update(updates)
cases_full.append(d)
tbl = pd.DataFrame(cases_full)
tbl = tbl.assign(continued=False)
for i, (_, ce) in enumerate(cases.items()):
n = ce.cx[lon_min:lon_max, lat_min:lat_max].mcs_id.nunique()
assert n in {1, 2}
if n == 1:
tbl.loc[i, "continued"] = True
tams.plot_tracked(ce, label="none", size=3.5, add_colorbar=True, cbar_kwargs=dict(fraction=0.05, shrink=0.3))
ax = plt.gca()
patch = mpl.patches.Polygon(
[(lon_min, lat_min), (lon_min, lat_max), (lon_max, lat_max), (lon_max, lat_min)],
ec="orangered",
fill=False,
transform=ccrs.PlateCarree(),
)
ax.add_patch(patch)
# proj = ccrs.PlateCarree()
# fig, axs = plt.subplots(m, n, figsize=(8, 5), sharex=True, sharey=True, subplot_kw=dict(projection=proj), constrained_layout=True)
# for i, ((key, ce), ax) in enumerate(zip(cases.items(), axs.flat)):
# if i == 0:
# assert key == "default"
# tams.plot_tracked(ce)
# tams.plot_tracked(ce.cx[lon_min:lon_max, lat_min:lat_max], ax=ax)
# ax.text(0.03, 0.98, key, ha="left", va="top", transform=ax.transAxes, size=8)
# gl = ax.gridlines(draw_labels=True, color="none")
# if not i < n:
# gl.top_labels = False
# if not i % n == 0:
# gl.left_labels = False
with open("sat-tracking-options-table.txt", "w") as f:
f.write(tbl.assign(continued=tbl.continued.map({True: "y", False: "n"})).style.to_latex())
plt.savefig("sat-tracking-options-ces-box.pdf", bbox_inches="tight", pad_inches=0.05, transparent=False)
For sample MPAS lat/lon dataset#
ds = tams.load_example_mpas().isel(time=slice(1, None)).rename_vars(tb="ctt", precip="pr")
ds
<xarray.Dataset>
Dimensions: (time: 126, lon: 341, lat: 180)
Coordinates:
* time (time) datetime64[ns] 2006-09-08T13:00:00 ... 2006-09-13T18:00:00
* lon (lon) float64 85.0 85.25 85.5 85.75 ... 169.2 169.5 169.8 170.0
* lat (lat) float64 -4.875 -4.625 -4.375 -4.125 ... 39.38 39.62 39.88
Data variables:
ctt (time, lat, lon) float32 266.8 266.9 267.0 ... 266.0 265.9 265.7
pr (time, lat, lon) float32 ...Identify CEs using different CTT thresholds#
For CEs (not necessarily for MCSs), the CTT threshold only affects identification.
Here we use tams.data_in_contours() to add a few cloud-top temperature
and precipitation rate statistics to our CE dataset.
%%time
ctt = ds.ctt
precip = ds.pr
thresh = 235
thresh_core = 219
cases_ces = {}
for delta in [-15, 0, 5]:
if delta == 0:
key = rf"default ($T = {thresh}\,\mathrm{{K}}$, $T_\mathrm{{core}} = {thresh_core}\,\mathrm{{K}}$)"
else:
s_delta = str(delta) if delta <= 0 else f"+{delta}"
key = rf"$(T, T_\mathrm{{core}}) {s_delta}$"
ces = tams.identify(ctt,
parallel=True,
ctt_threshold=thresh + delta,
ctt_core_threshold=thresh_core + delta,
)[0]
assert len(ces) == ds.sizes["time"]
# Add min Tb and avg precip -- parallel
# NOTE: parallel attempt giving can't pickle threadlock error
def fun(ds, ce):
ce = tams.data_in_contours(ds.ctt, ce, agg=("mean", "min",), merge=True)
ce = tams.data_in_contours(ds.pr, ce, agg=("mean",), merge=True)
return ce
ces_ = Parallel(n_jobs=-2, verbose=10)(
delayed(fun)(ds.isel(time=i).copy(deep=False), ce.copy())
for i, ce in enumerate(ces)
)
cases_ces[key] = ces_
# # Add min Tb and avg precip -- serial
# for i, ce in enumerate(ces):
# ce = tams.data_in_contours(ctt.isel(time=i), ce, agg=("min",), merge=True)
# ce = tams.data_in_contours(precip.isel(time=i), ce, agg=("mean",), merge=True)
# ces[i] = ce
# print(".", end="")
# print()
# cases_ces[key] = ces
Show code cell output
[Parallel(n_jobs=-2)]: Using backend LokyBackend with 11 concurrent workers.
[Parallel(n_jobs=-2)]: Done 3 tasks | elapsed: 0.8s
[Parallel(n_jobs=-2)]: Done 10 tasks | elapsed: 1.8s
[Parallel(n_jobs=-2)]: Done 19 tasks | elapsed: 3.6s
[Parallel(n_jobs=-2)]: Done 28 tasks | elapsed: 4.5s
[Parallel(n_jobs=-2)]: Done 39 tasks | elapsed: 5.2s
[Parallel(n_jobs=-2)]: Done 50 tasks | elapsed: 6.2s
[Parallel(n_jobs=-2)]: Done 63 tasks | elapsed: 7.2s
[Parallel(n_jobs=-2)]: Done 76 tasks | elapsed: 8.5s
[Parallel(n_jobs=-2)]: Done 91 tasks | elapsed: 9.6s
[Parallel(n_jobs=-2)]: Done 118 out of 126 | elapsed: 11.6s remaining: 0.7s
[Parallel(n_jobs=-2)]: Done 126 out of 126 | elapsed: 12.2s finished
[Parallel(n_jobs=-2)]: Using backend LokyBackend with 11 concurrent workers.
[Parallel(n_jobs=-2)]: Done 3 tasks | elapsed: 1.3s
[Parallel(n_jobs=-2)]: Done 10 tasks | elapsed: 1.4s
[Parallel(n_jobs=-2)]: Done 19 tasks | elapsed: 2.6s
[Parallel(n_jobs=-2)]: Done 28 tasks | elapsed: 3.7s
[Parallel(n_jobs=-2)]: Done 39 tasks | elapsed: 4.9s
[Parallel(n_jobs=-2)]: Done 50 tasks | elapsed: 6.0s
[Parallel(n_jobs=-2)]: Done 63 tasks | elapsed: 7.3s
[Parallel(n_jobs=-2)]: Done 76 tasks | elapsed: 8.6s
[Parallel(n_jobs=-2)]: Done 91 tasks | elapsed: 10.6s
[Parallel(n_jobs=-2)]: Done 118 out of 126 | elapsed: 13.5s remaining: 0.8s
[Parallel(n_jobs=-2)]: Done 126 out of 126 | elapsed: 14.2s finished
[Parallel(n_jobs=-2)]: Using backend LokyBackend with 11 concurrent workers.
[Parallel(n_jobs=-2)]: Done 3 tasks | elapsed: 1.1s
[Parallel(n_jobs=-2)]: Done 10 tasks | elapsed: 1.3s
[Parallel(n_jobs=-2)]: Done 19 tasks | elapsed: 2.8s
[Parallel(n_jobs=-2)]: Done 28 tasks | elapsed: 4.3s
[Parallel(n_jobs=-2)]: Done 39 tasks | elapsed: 5.9s
[Parallel(n_jobs=-2)]: Done 50 tasks | elapsed: 7.6s
[Parallel(n_jobs=-2)]: Done 63 tasks | elapsed: 9.5s
[Parallel(n_jobs=-2)]: Done 76 tasks | elapsed: 11.5s
[Parallel(n_jobs=-2)]: Done 91 tasks | elapsed: 13.8s
[Parallel(n_jobs=-2)]: Done 118 out of 126 | elapsed: 17.7s remaining: 1.1s
[Parallel(n_jobs=-2)]: Done 126 out of 126 | elapsed: 18.4s finished
[Parallel(n_jobs=-2)]: Using backend LokyBackend with 11 concurrent workers.
[Parallel(n_jobs=-2)]: Done 3 tasks | elapsed: 1.6s
[Parallel(n_jobs=-2)]: Done 10 tasks | elapsed: 1.9s
[Parallel(n_jobs=-2)]: Done 19 tasks | elapsed: 3.6s
[Parallel(n_jobs=-2)]: Done 28 tasks | elapsed: 4.9s
[Parallel(n_jobs=-2)]: Done 39 tasks | elapsed: 6.3s
[Parallel(n_jobs=-2)]: Done 50 tasks | elapsed: 8.1s
[Parallel(n_jobs=-2)]: Done 63 tasks | elapsed: 9.8s
[Parallel(n_jobs=-2)]: Done 76 tasks | elapsed: 11.7s
[Parallel(n_jobs=-2)]: Done 91 tasks | elapsed: 14.6s
[Parallel(n_jobs=-2)]: Done 118 out of 126 | elapsed: 18.2s remaining: 1.1s
[Parallel(n_jobs=-2)]: Done 126 out of 126 | elapsed: 19.2s finished
[Parallel(n_jobs=-2)]: Using backend LokyBackend with 11 concurrent workers.
[Parallel(n_jobs=-2)]: Done 3 tasks | elapsed: 1.2s
[Parallel(n_jobs=-2)]: Done 10 tasks | elapsed: 1.3s
[Parallel(n_jobs=-2)]: Done 19 tasks | elapsed: 2.9s
[Parallel(n_jobs=-2)]: Done 28 tasks | elapsed: 4.7s
[Parallel(n_jobs=-2)]: Done 39 tasks | elapsed: 6.4s
[Parallel(n_jobs=-2)]: Done 50 tasks | elapsed: 8.3s
[Parallel(n_jobs=-2)]: Done 63 tasks | elapsed: 10.3s
[Parallel(n_jobs=-2)]: Done 76 tasks | elapsed: 12.3s
[Parallel(n_jobs=-2)]: Done 91 tasks | elapsed: 14.5s
[Parallel(n_jobs=-2)]: Done 118 out of 126 | elapsed: 18.9s remaining: 1.2s
[Parallel(n_jobs=-2)]: Done 126 out of 126 | elapsed: 19.8s finished
[Parallel(n_jobs=-2)]: Using backend LokyBackend with 11 concurrent workers.
[Parallel(n_jobs=-2)]: Done 3 tasks | elapsed: 1.9s
[Parallel(n_jobs=-2)]: Done 10 tasks | elapsed: 2.1s
[Parallel(n_jobs=-2)]: Done 19 tasks | elapsed: 3.8s
[Parallel(n_jobs=-2)]: Done 28 tasks | elapsed: 5.3s
[Parallel(n_jobs=-2)]: Done 39 tasks | elapsed: 7.1s
[Parallel(n_jobs=-2)]: Done 50 tasks | elapsed: 8.9s
[Parallel(n_jobs=-2)]: Done 63 tasks | elapsed: 10.5s
[Parallel(n_jobs=-2)]: Done 76 tasks | elapsed: 12.7s
[Parallel(n_jobs=-2)]: Done 91 tasks | elapsed: 15.2s
[Parallel(n_jobs=-2)]: Done 118 out of 126 | elapsed: 19.3s remaining: 1.2s
CPU times: total: 7.64 s
Wall time: 1min 44s
[Parallel(n_jobs=-2)]: Done 126 out of 126 | elapsed: 20.2s finished
Track for different \(u\) projection values#
For each of our CTT threshold cases, we track using different \(u\) projection values, bringing the case count to 9.
%%time
cases = {}
time = ctt.time.values.tolist()
projs = [-5, -10, -12]
for proj in projs:
for thresh_key, ces in cases_ces.items():
key = rf"{thresh_key}, $u_\mathrm{{proj}} = {proj}\,\mathrm{{m}}\,\mathrm{{s}}^{{-1}}$"
cases[key] = tams.track(ces, time, u_projection=proj)
CPU times: total: 1min 40s
Wall time: 1min 40s
Check the case labels#
fig, ax = plt.subplots(figsize=(4, 3))
ax.axis("off")
lines = []
for i, k in enumerate(cases):
lines.append(f"{i+1}. `{k}`")
y = len(cases) - i - 0.5
ax.text(0, y, i + 1, va="center", ha="left", color="orangered")
ax.text(0.07, y, k, va="center", ha="left")
ax.set_ylim(ymax=len(cases))
display(Markdown("\n\n".join(lines)))
$(T, T_\mathrm{core}) -15$, $u_\mathrm{proj} = -5\,\mathrm{m}\,\mathrm{s}^{-1}$default ($T = 235\,\mathrm{K}$, $T_\mathrm{core} = 219\,\mathrm{K}$), $u_\mathrm{proj} = -5\,\mathrm{m}\,\mathrm{s}^{-1}$$(T, T_\mathrm{core}) +5$, $u_\mathrm{proj} = -5\,\mathrm{m}\,\mathrm{s}^{-1}$$(T, T_\mathrm{core}) -15$, $u_\mathrm{proj} = -10\,\mathrm{m}\,\mathrm{s}^{-1}$default ($T = 235\,\mathrm{K}$, $T_\mathrm{core} = 219\,\mathrm{K}$), $u_\mathrm{proj} = -10\,\mathrm{m}\,\mathrm{s}^{-1}$$(T, T_\mathrm{core}) +5$, $u_\mathrm{proj} = -10\,\mathrm{m}\,\mathrm{s}^{-1}$$(T, T_\mathrm{core}) -15$, $u_\mathrm{proj} = -12\,\mathrm{m}\,\mathrm{s}^{-1}$default ($T = 235\,\mathrm{K}$, $T_\mathrm{core} = 219\,\mathrm{K}$), $u_\mathrm{proj} = -12\,\mathrm{m}\,\mathrm{s}^{-1}$$(T, T_\mathrm{core}) +5$, $u_\mathrm{proj} = -12\,\mathrm{m}\,\mathrm{s}^{-1}$
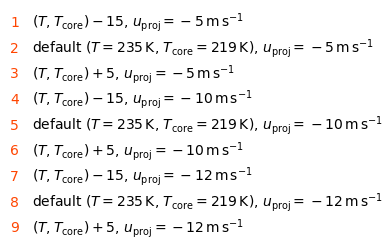
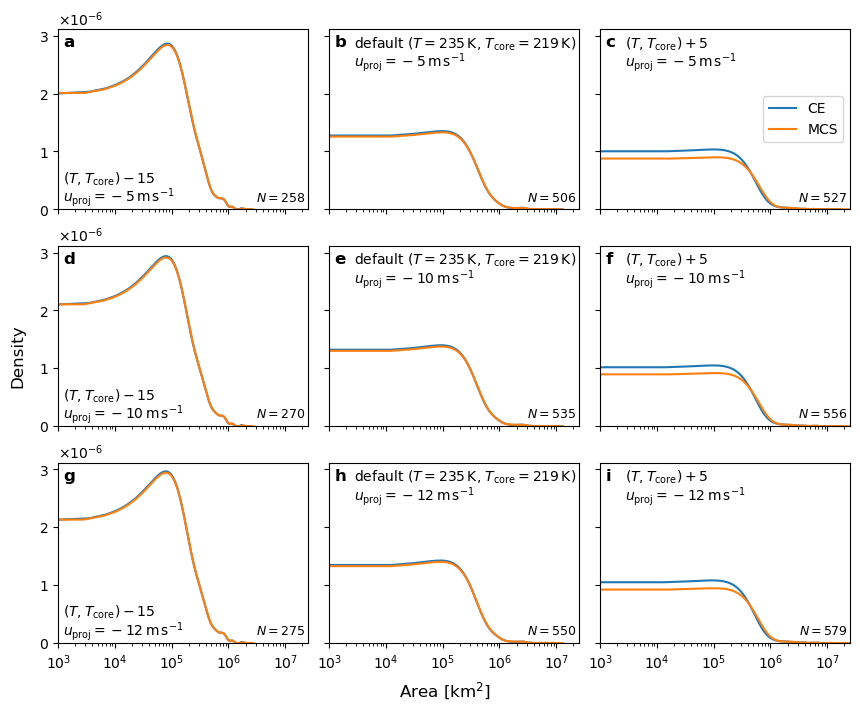
Plot area dist#
m = len(projs)
n = 3
assert m * n == len(cases)
fig, axs = plt.subplots(m, n, figsize=(8.2, 7), sharex=True, sharey=True, constrained_layout=True)
for a, (key, ce), ax in zip(ascii_lowercase, cases.items(), axs.flat):
n = ce.mcs_id.nunique()
# Dist of max area
x = ce.groupby("mcs_id").area_km2.max()
x.plot.kde(ax=ax, label="CE")
mcs = ce.groupby(["mcs_id", "itime"]).area_km2.sum()
x2 = mcs.groupby("mcs_id").max()
x2.plot.kde(ax=ax, label="MCS")
# ax.text(0.99, 0.98, f"$N={n}$", size=9, ha="right", va="top", transform=ax.transAxes)
ax.text(0.99, 0.03, f"$N={n}$", size=9, ha="right", va="bottom", transform=ax.transAxes)
# ax.set_title(key, size=9)
i = key.index(", $u_")
l1, l2 = key[:i], key[i+2:]
if ax.get_subplotspec().is_first_col():
xt, yt = 0.02, 0.22
else:
xt, yt = 0.1, 0.97
ax.text(xt, yt, l1, size=10, ha="left", va="top", transform=ax.transAxes)
ax.text(xt, yt - 0.09, l2, size=10, ha="left", va="top", transform=ax.transAxes)
ax.set_ylabel("")
ax.text(0.02, 0.97, f"{a}",
size=12, weight="bold",
ha="left", va="top",
transform=ax.transAxes,
)
if a == "c":
ax.legend(loc="center right")
fig.supxlabel("Area [km$^2$]")
fig.supylabel("Density", x=-0.03)
# ax.set_xlim(xmin=0)
ax.set_xlim(xmin=1000); ax.set_xscale("log")
ax.set_ylim(ymin=0)
fig.savefig("mpas-tracking-options-area-kde.pdf", bbox_inches="tight", pad_inches=0.05, transparent=False)
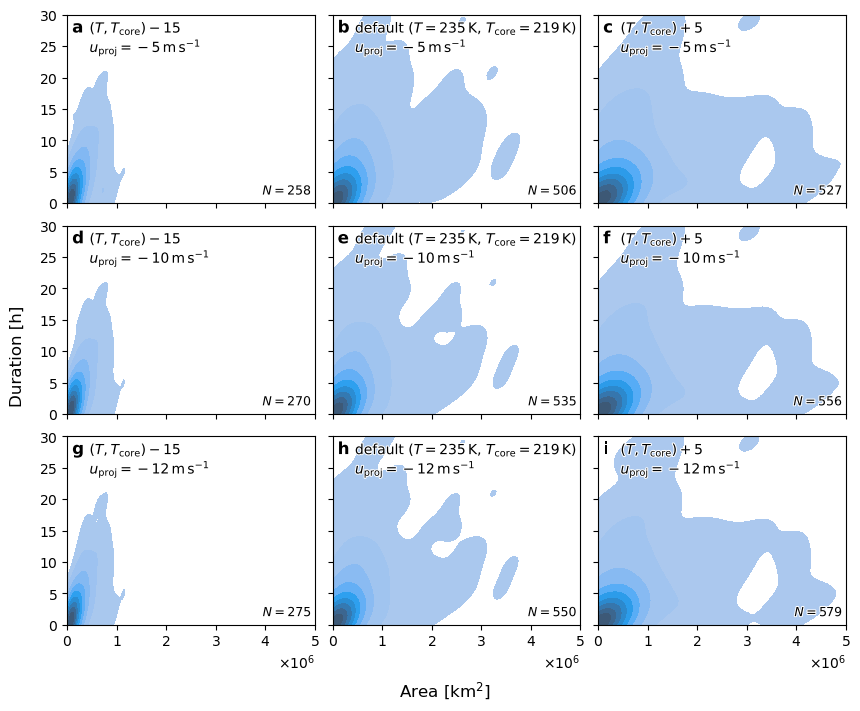
Plot 2-D duration and area dist#
m = len(projs)
n = 3
assert m * n == len(cases)
fig, axs = plt.subplots(m, n, figsize=(8.2, 7), sharex=True, sharey=True, constrained_layout=True)
for a, (key, ce), ax in zip(ascii_lowercase, cases.items(), axs.flat):
n = ce.mcs_id.nunique()
gb = ce.groupby("mcs_id")
area = gb.area_km2.max()
dur = ((gb.time.max() - gb.time.min()).dt.total_seconds() / 3600).rename("duration_h")
data = pd.concat([area, dur], axis="columns")
# sns.jointplot(x="area_km2", y="duration_h", kind="kde", joint_kws=dict(fill=True), ax=ax, data=data)
sns.kdeplot(x="area_km2", y="duration_h",
fill=True,
common_norm=True, clip=(0, None), hue_norm=(0, 0.5),
ax=ax, data=data,
)
# TODO: ensure same levels?
# sns.rugplot(x="area_km2", y="duration_h", ax=ax, data=data)
#ax.text(0.99, 0.98, f"$N={n}$", size=9, ha="right", va="top", transform=ax.transAxes)
ax.text(0.99, 0.03, f"$N={n}$",
size=9,
ha="right", va="bottom",
path_effects=[path_effects.withStroke(linewidth=1.5, foreground="w")],
transform=ax.transAxes,
)
# ax.set_title(key, size=10)
i = key.index(", $u_")
l1, l2 = key[:i], key[i+2:]
xt, yt = 0.09, 0.975
kws = dict(
size=10,
ha="left", va="top",
path_effects=[path_effects.withStroke(linewidth=1.5, foreground="w")],
transform=ax.transAxes,
)
ax.text(xt, yt, l1, **kws)
ax.text(xt, yt - 0.09, l2, **kws)
ax.set_xlabel("")
ax.set_ylabel("")
ax.text(0.02, 0.97, f"{a}",
size=12, weight="bold",
ha="left", va="top",
path_effects=[path_effects.withStroke(linewidth=1.5, foreground="w")],
transform=ax.transAxes,
)
fig.supxlabel("Area [km$^2$]")
fig.supylabel("Duration [h]", x=-0.03)
ax.set_xlim(xmin=0, xmax=0.5e7)
ax.set_ylim(ymin=0, ymax=30)
fig.savefig("mpas-tracking-options-duration-area-2d-kde.pdf", bbox_inches="tight", pad_inches=0.05, transparent=False)
list(cases.values())[0].head(3)
| geometry | area_km2 | inds219 | area219_km2 | cs219 | mean_ctt | min_ctt | mean_pr | time | itime | dtime | mcs_id | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | POLYGON ((152.50000 8.93176, 154.09671 11.1250... | 65189.730096 | [1, 2, 3, 4] | 7632.145729 | MULTIPOLYGON (((154.00000 11.33169, 154.00332 ... | 223.781916 | 199.467636 | 2.174953 | 2006-09-08 13:00:00 | 0 | 0 days 01:00:00 | 0 |
| 1 | POLYGON ((105.00000 20.86816, 105.04808 20.875... | 43468.589233 | [8] | 11043.783834 | MULTIPOLYGON (((105.00000 21.04446, 105.25000 ... | 217.004782 | 196.3992 | 3.237674 | 2006-09-08 13:00:00 | 0 | 0 days 01:00:00 | 1 |
| 2 | POLYGON ((103.75000 0.07912, 103.83888 0.12500... | 98642.412303 | [9, 10, 11] | 38699.492801 | MULTIPOLYGON (((103.25000 1.66696, 103.28213 1... | 216.819586 | 195.909149 | 3.098599 | 2006-09-08 13:00:00 | 0 | 0 days 01:00:00 | 2 |
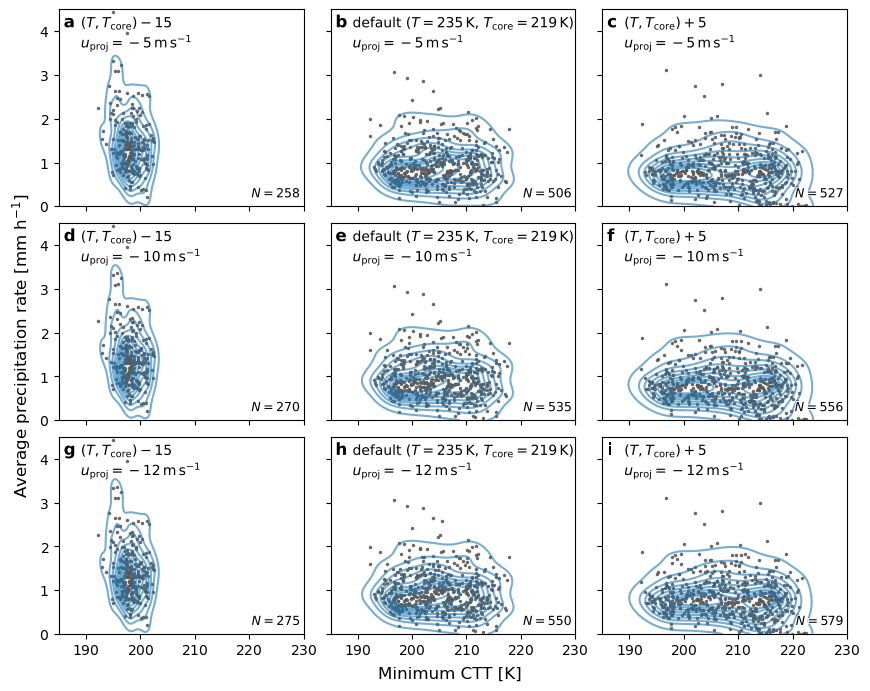
Plot 2-D CTT and precip dist#
m = len(projs)
n = 3
assert m * n == len(cases)
fig, axs = plt.subplots(m, n, figsize=(8.3, 6.8), sharex=True, sharey=True, constrained_layout=True)
for a, (key, ce), ax in zip(ascii_lowercase, cases.items(), axs.flat):
n = ce.mcs_id.nunique()
gb = ce.groupby("mcs_id")
tb = gb.min_ctt.min().astype(float) # np.sqrt not supported for `Float64`
pr = gb.mean_pr.mean().astype(float) # TODO: area-weighted average of CE precip at each time?
data = pd.concat([tb, pr], axis="columns")
ax.plot(tb, pr, ".", ms=5, mec="none", mfc="0.35", alpha=0.9)
sns.kdeplot(x="min_ctt", y="mean_pr",
fill=False, color=sns.color_palette()[0], alpha=0.6,
common_norm=True, clip=(0, None),
ax=ax, data=data,
)
ax.text(0.99, 0.03, f"$N={n}$",
size=9,
ha="right", va="bottom",
path_effects=[path_effects.withStroke(linewidth=1.5, foreground="w")],
transform=ax.transAxes,
)
i = key.index(", $u_")
l1, l2 = key[:i], key[i+2:]
xt, yt = 0.09, 0.975
kws = dict(
size=10,
ha="left", va="top",
path_effects=[path_effects.withStroke(linewidth=1.5, foreground="w")],
transform=ax.transAxes,
)
ax.text(xt, yt, l1, **kws)
ax.text(xt, yt - 0.09, l2, **kws)
ax.set_xlabel("")
ax.set_ylabel("")
ax.text(0.02, 0.97, f"{a}",
size=12, weight="bold",
ha="left", va="top",
path_effects=[path_effects.withStroke(linewidth=1.5, foreground="w")],
transform=ax.transAxes,
)
fig.supxlabel("Minimum CTT [K]")
fig.supylabel("Average precipitation rate [mm h$^{-1}$]", x=-0.03)
ax.set_xlim(xmin=185, xmax=230)
ax.set_ylim(ymin=0, ymax=4.5)
fig.savefig("mpas-tracking-options-tb-pr-2d-kde.pdf", bbox_inches="tight", pad_inches=0.05, transparent=False)